In the life sciences, protein quantification is a frequently used research technique. Many researchers have faced this common dilemma: with more than a dozen protein quantification methods available—ELISA, BCA, chemiluminescence, and more—which one should you choose?

Reed Biotech presents a three-part series on protein quantification. This first installment serves as the basics, breaking down the core classification logic behind the main methods and providing a detailed explanation of non-specific protein detection and common single-factor specific detection. From principles and practical operation to suitable applications, this guide gives a one-stop overview of fundamental methods, helping you quickly master routine protein quantification experiments.
I. Two Core Categories
Protein quantification methods are primarily classified based on whether they rely on antigen-antibody specific binding. This gives rise to two main categories, each suited to different experimental needs, making the logic clear and straightforward.
1. Non-specific Protein Quantification (Quick Total Protein Measurement)
Core Features:
· Does not rely on antigen-antibody binding.
· Quantifies proteins based on common features (e.g., UV absorption of aromatic amino acids or dye-binding color reactions).
· Measures total protein content only.
· Suitable when you only need total protein concentration without distinguishing specific proteins.
This article introduces three commonly used methods.
2. Specific Protein Quantification (Targeted Protein Measurement)
Core Features:
· Requires antigen-antibody specific binding.
· Quantifies a target protein with high specificity.
· Widely used in research and clinical applications.
· Suitable for precise measurement of known target proteins.
This article covers three basic single-factor detection methods. Advanced topics like ultra-sensitive and multiplex detection will be discussed in later installments.
II. Non-specific Total Protein Detection: 3 Methods
1. UV Absorption
Principle:
Aromatic amino acids in proteins (e.g., tyrosine, tryptophan) absorb UV light at 280 nm. Within a certain concentration range, absorbance is proportional to protein concentration. No color reagents are required.
Procedure:
· Dilute sample to appropriate concentration → place in cuvette → set UV spectrophotometer to 280 nm (use blank buffer as control) → measure absorbance → calculate concentration.
· Time: 5–10 minutes.
Advantages:
· Simple, fast, low-cost, non-destructive (sample can be reused).
Considerations:
· Low specificity; may be affected by nucleic acids (strong 260 nm absorption overlaps at 280 nm). Correct using A280/A260 ratio if nucleic acids are present.
Applications:
· High-purity protein samples, e.g., purified protein concentration measurement, monitoring protein purification processes.
2. BCA Assay
Principle:
Proteins reduce Cu²⁺ to Cu⁺ under alkaline conditions. Cu⁺ reacts with BCA reagent to form a purple complex, with maximum absorbance at 562 nm. Absorbance correlates with protein concentration within a specific range.
Procedure:
· Prepare BCA working solution → add standards/samples → add BCA solution → incubate 37°C for 30 min → read absorbance at 562 nm.
Advantages:
· Wide linear range, good resistance to interference (tolerates some detergents), stable results. Often the preferred method for total protein quantification.
Applications:
· Total protein measurement in various sample types, high-throughput detection.
3. Coomassie Brilliant Blue (Bradford) Assay
G-250 (red color)+ Protein —— Measure absorbance at 595 nm (blue)
Principle:
Coomassie Brilliant Blue G-250 dye is red in acidic solution. Upon binding proteins, it turns blue, with maximum absorbance at 595 nm. Absorbance correlates with protein concentration.
Procedure:
· Prepare dye solution → add standards/samples → add dye → incubate at room temperature 5–10 min → measure absorbance at 595 nm.
Advantages:
· Low cost, simple, suitable for rapid large-scale screening.
Considerations:
· Narrow linear range, sensitive to detergents and electrolytes.
Applications:
· High-purity protein samples, monitoring protein purification processes.
III. Specific Single-Factor Detection: ELISA, Chemiluminescence, TR-FRET
1. ELISA (Enzyme-Linked Immunosorbent Assay)
Principle:
Antigens or antibodies are coated on polystyrene plates. Through antigen-antibody specific binding, enzyme conjugates catalyze substrate color change. Color intensity correlates with protein concentration.
Procedure (sandwich ELISA example):
· Coat capture antibody → add standards/samples → add biotinylated detection antibody → add enzyme conjugate → develop color → read on plate reader.
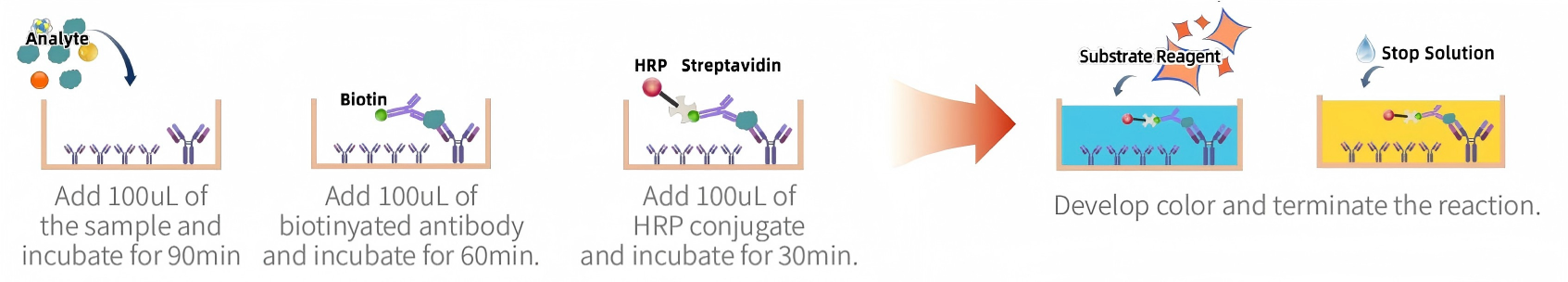
Advantages:
· Mature technique, simple, compatible with serum, plasma, and cell culture supernatants.
Applications:
· Routine research protein quantification, beginner-friendly experiments.
2. Chemiluminescence (CLIA)
Principle:
Based on antigen-antibody binding, chemiluminescent substrates generate visible light. Signal intensity correlates with target protein concentration. No external light source is needed, background is lower than traditional colorimetric methods, and sensitivity is higher than ELISA.
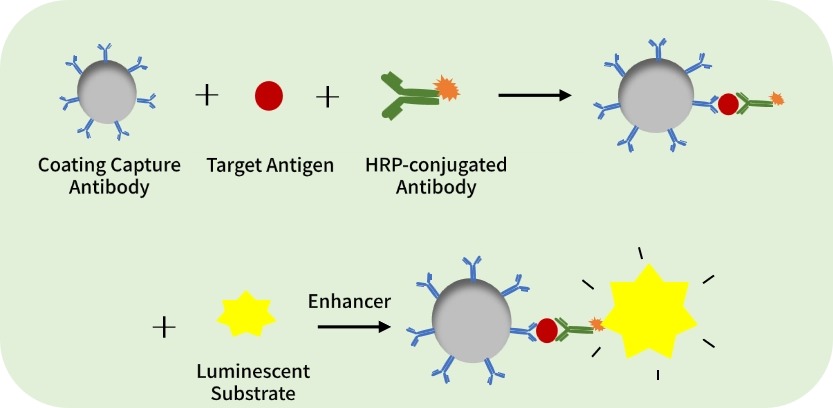
Procedure:
· Coat antibody → add sample → add enzyme-labeled antibody → add luminescent substrate → measure and calculate concentration.
Advantages:
· High sensitivity, fast, moderately priced, suitable for experiments requiring better sensitivity without high cost.
Applications:
· Clinical diagnostics, research where ELISA sensitivity is insufficient, high-sensitivity routine protein quantification.
3. TR-FRET (Time-Resolved Fluorescence Resonance Energy Transfer)
Principle:
FRET occurs between donor and acceptor fluorophores conjugated to specific antibodies. When bound to target protein (<10 nm distance), energy transfer generates quantifiable fluorescence.
Procedure:
· Mix sample with donor/acceptor antibodies → incubate in same system → measure with time-resolved fluorescence reader.
Advantages:
· High signal-to-noise ratio, homogeneous (no washing), fast, simple.
Applications:
· Drug screening, protein interaction studies, rapid high-throughput protein quantification.
Considerations:
· Requires labeled antibodies; reaction conditions (temperature, time) are critical.
IV. Summary of Basics
1. Protein quantification is divided into two categories: specific detection (measure a single protein precisely) and non-specific detection (measure total protein).
2. Total protein measurement:
o Stability → BCA
o Speed → UV absorption
o Cost-saving → Bradford
3. Routine single-factor detection:
o ELISA → low-cost choice
o Chemiluminescence → high sensitivity alternative
o TR-FRET → no-wash, high-throughput option
Mastering these basics allows you to handle 80% of routine experiments.
In the next part of this series, we will cover advanced methods for “simultaneous detection of multiple proteins” and “precise quantification of low-abundance proteins,” such as Luminex and Simoa. Stay tuned!
